J. Andrew Bangham (1947 – 2014): Enterprising scientist who broke new ground in computational biology and image analysis
Andrew Bangham, former Professor in Computer Science at the University of East Anglia, Norwich, was a pioneering researcher with the rare ability to integrate diverse disciplines: from computer science and electrophysiology to plant development and visual art. He is best known for developing the theory of ‘sieves’, which find frequent application in modern computer vision including artistic rendering of images, and for interdisciplinary approaches to modelling plant growth and development. He was also an inspirational teacher who cultivated and encouraged talent during the birth and flowering of computer science.
Andrew was born just after the war into a scientific and artistic family. He was the eldest son of medical doctors Rosalind and Alec Bangham, and grew up outside Cambridge where Alec (later FRS) was a researcher at the Babraham Institute of Animal Physiology. Andrew’s village upbringing was filled with sailing, photography and painting; his uncle Patrick Heron had a strong influence on Andrew’s later interest in the perception of colour.
Andrew found his vocation soon after finishing high school. Unsure whether he would have the sufficient grades to study medicine, his father arranged for him to spend six months in the US, working with eminent physiologist Dan Tosteson at Duke University in North Carolina. He took to experimental research like a duck to water. Fired up by his experiences at Duke, and by the publication of his first paper, Andrew arrived at University College London (UCL) to study physiology.
The late 1960s and early 1970s were a tremendously exciting time for electrophysiology and UCL had multiple Nobel prize-winning physiologists working there. After his undergraduate degree—during which he met his future wife Kate, a medical student—he stayed on at UCL to complete a PhD in the Biophysics Department, where he worked on the electrophysiology of mitochondria. Further developing the lipid film techniques first acquired in the Tosteson lab, Andrew shared his work with colleagues at Cambridge and at the University of East Anglia (UEA).This led to an invitation to join the UEA Biology Department in 1973, in order to continue studying the electrical properties of cells and tissues.
Computational methods were still in their infancy, and Andrew became one of the first to use computers to address physiological questions. He wrote computer programmes to interpret the very noisy electrical traces produced by his experiments, inventing new filtering techniques to pick out signals in the noise. In the 1970s, the challenges of using computing in research were not purely technical, but also financial. Andrew established new ways of raising research money, setting up contracts to do modelling for local commercial companies on the University’s mainframe computers.
Through this work he was eventually able to buy a state-of-the-art Sun computer, which offered enough computing power for work on a new class of information filters, which he would later call ‘sieves’. By analogy with physical sieves, which pass objects depending on their size, computational sieves filter signals using medians—in contrast to the more conventional mean-based filters. Andrew realized that the potential for sieves was in the growing field of computer vision.
Eventually he decided to devote himself full-time to computational problems, moving in mid-1980s from UEA’s Biology Department to its School of Information Systems. Undaunted by the need to master a whole new field of mathematics known as mathematical morphology, Andrew formulated a new theoretical treatment for computer vision. He extended his funding approach to his new department by setting up contacts between numerous research groups and local and national companies, such as the local Norwich shoe company Start-rite, Britain’s Post Office, and the Independent Television Commission.
Andrew also realised that his median sieves allowed him to extract striking colour patterns from photographs, reminiscent of the paintings by Heron. Andrew became deeply interested in the relationship between median sieves and the ways in which human brains interpret visual signals. He developed software—later known as PhotoArtmaster—that offered tools for the digital alteration and analysis of photographs, and which he sold through his company Fo2Pix.
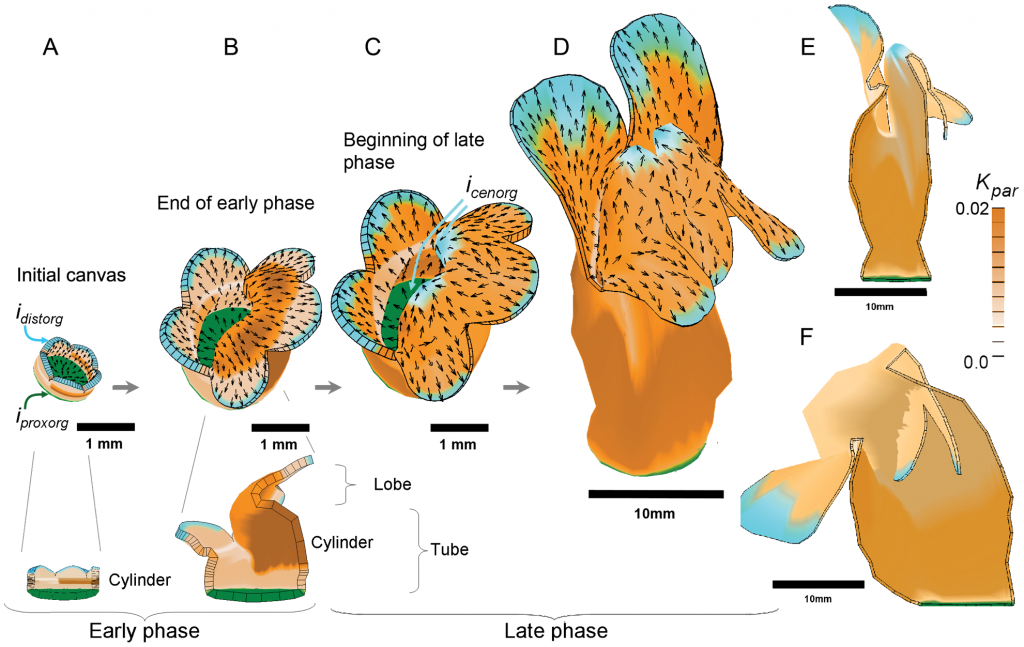
Throughout this period of entrepreneurship and programming, Andrew retained a strong interest in biology. In the mid-1990s he began collaborating with Professor Enrico Coen at the John Innes Centre, who shared Andrew’s passion for understanding pattern and form. They tackled the problem of how the petals of the garden snapdragon grow using a combination of computational and experimental approaches. Inspired by these experiences, Andrew established UEA’s D’Arcy Thompson Centre for Computational Biology.
A passionate sailor, and devoted to his family, Andrew had a broad network of friends, colleagues and students in Norwich and beyond. At a celebration of his scientific work at the John Innes Centre, speaker after speaker talked of Andrew’s influential role in their work, of his enthusiasm, and of his ability to articulate the links between individual problems and whole scientific fields. Colleagues described him as the unusual combination of unconventional thinker, wonderful host, inspiring teacher and supportive colleague.
Andrew, Enrico and colleagues published their work on the genetic control of tissue growth in PLOS Computational Biology and PLOS Biology:
http://journals.plos.org/ploscompbiol/article?id=10.1371/journal.pcbi.1002071
http://journals.plos.org/plosbiology/article?id=10.1371/journal.pbio.1000537


[…] (adsbygoogle = window.adsbygoogle || []).push({}); adUnit = document.getElementById("google-ads-E3RH"); adWidth = adUnit.offsetWidth; if ( adWidth >= 999999 ) { /* GETTING THE FIRST IF OUT OF THE WAY */ } else if ( adWidth >= 728 ) { document.getElementById("GARDasync_E3RH").innerHTML = ""; } else if ( adWidth >= 468 ) { document.getElementById("GARDasync_E3RH").innerHTML = ""; } else if ( adWidth >= 250 ) { document.getElementById("GARDasync_E3RH").innerHTML = ""; } Read more: https://blogs.plos.org/biologue/2015/07/08/j-andrew-bangham-1947-2014-enterprising-scientist-who-brok… […]
[…] PLOS Blogs: J. Andrew Bangham (1947–2014): Enterprising scientist who broke new ground in computational biolog… […]
[…] In a previous blog I mentioned about a work colleague who died from prostate cancer last summer. One of his students has now produced a detailed account of his pioneering work (https://blogs.plos.org/biologue/2015/07/08/j-andrew-bangham-1947-2014-enterprising-scientist-who-brok…). […]