Understanding Images: Coat of Invisibility – How Trypanosoma brucei Evades Recognition
Author: Galadriel Hovel-Miner, George Washington University, Washington DC, United States.
Competing Interests: Galadriel Hovel-Miner is an author of the article discussed in this blog.
Image Credit: Mark Field, University of Cambridge
Every pathogen faces a similar predicament of how to exploit nutrient-rich, but often inhospitable, host environments. The cellular surface is the primary interface of the host-pathogen interaction, where receptors of immunity reside, ready to detect and respond to foreign molecules exposed on the surface of viruses, bacteria, and eukaryotic pathogens. Commonly, pathogens escape host immunity by hiding (i.e. inhabiting a site ‘unseen’ by the immune system) or by hiding in plain sight through the variation of their cell surface molecules (i.e. antigenic variation). Natural pressure from the immune system has resulted in the convergent evolution of antigenic variation among both viruses and diverse phyla of microbes. These systems share fascinating genetic characteristics that include variant multigene families, hypermutation, recombination-based gene rearrangements, and subtelomeric organization of variant genes.
Infection with Trypanosoma brucei

The genetic features and process of antigenic variation itself are exemplified by unicellular African trypanosome parasites (see Issue Image). Following transmission to the bloodstream by the bite of a tsetse fly, Trypanosoma brucei and its other African cousins are able to grow robustly while their primary surface antigen elicits a strong antibody-mediated immune response. Yet before the immune system can clear this aggressive infection of the blood, subpopulations of (or individual) parasites switch their protein coats, rendering the new variants ‘invisible’ to the existing antibodies.

The result is a chronic infection, during which parasites find their way into the central nervous system and, ultimately, the brain. These infections cause African sleeping sickness in humans, an illness resulting in death if left untreated, and nagana in livestock, which is estimated to cost US$1-3 billion per year in lost domestic produce contributing to worldwide poverty.
Variant Surface Glycoprotein is Paramount to T. brucei’s Adaptation to the Bloodstream
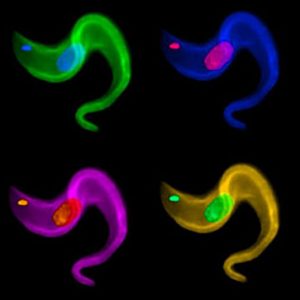
Central to T. brucei’s bloodstream adaptation is the variant surface glycoprotein (VSG) that forms its antigenic coat. In the featured image for the May issue of PLOS Genetics, we show the unicellular parasite stained for DNA, highlighting both the nucleus and kinetoplast (a defining organelle of these protozoa), and with an antibody against a single VSG coat protein. The differential color enhancements beautifully illustrate the parasite’s ability to switch the expression of its entire protein coat from one VSG coat to the next. VSG-encoding genes comprise a multigene family consisting of more than 2000 genes and pseudogenes distributed throughout the genome. Yet only one VSG gene can be expressed at a given time from a specialized site near the end of a chromosome. Sites of VSG expression also contain a long region of repetitive DNA in very close proximity to the VSG gene. Interestingly, shorter iterations of the same repeat sequence are found near most VSGs throughout the genome. To switch from one active VSG to the next usually requires that a silent gene is copied into the actively expressed site by specific, but as yet poorly characterized, DNA recombination events.
DNA Repeats Enhance Selection of VSGs in Time with the Cell Cycle
In May’s PLOS Genetics issue image article we report our findings defining a role of the VSG proximal DNA repeats in T. brucei antigenic variation. Throughout the living world the properties of repetitive DNA elements have been adapted to carry out specific activities. They can function as sites of protein binding, provide homology for recombination, stabilize chromosome ends (as seen with telomeric repeats), or lead to instability that results in genetic disorders (such as Huntington’s disease and Fragile X syndrome in humans). In this article we genetically manipulated the repeat regions in the VSG-expressing site to determine their effects on VSG switching.

We demonstrated that the repeats are required for the selection of VSGs throughout the genome during coat switching and define a discrete, sequence-specific element that is able to carry out this function. Two intriguing findings to arise from this study are that the DNA repeats enhance selection of VSGs from diverse genomic locations and that the process appears to be linked to the timing of the cell cycle. Limiting the selection of a sufficiently diverse VSG repertoire would likely result in parasites with a diminished capacity for chronic host infections.
Battle of Adaptation
African trypanosomes belong to an ancient clade and it is likely that they coevolved alongside their mammalian hosts. As the host immune system developed its astonishing capacity for antibody diversity and specificity, trypanosomes evolved a system of genetic variation to efficiently escape it. Both systems (immunity and antigenic variation) rely on similar genetic themes: repetitive DNA elements, variant multigene families, and generation of diversity through recombination. When considering the known components of each system, one could easily argue that the host’s defenses are more elaborate and elegant when compared to the parasite’s offensive strategies. Sadly, in this head-to-head battle of adaptation, the parasite will ultimately win.
Reference
